HIPC scRNA-seq Annotation Benchmark Challenge
Welcome to the HIPC scRNA-seq Annotation Benchmark. This initiative is designed to evaluate the consistency, accuracy, and granularity of cell-type annotations across specialized bioinformatics teams. Using high-quality immune cell datasets, we aim to establish a gold standard for single-cell annotation for immune response datasets. We are accepting entries throughout the challenge period. If you are interested in joining this benchmark challenge, please sign up here.
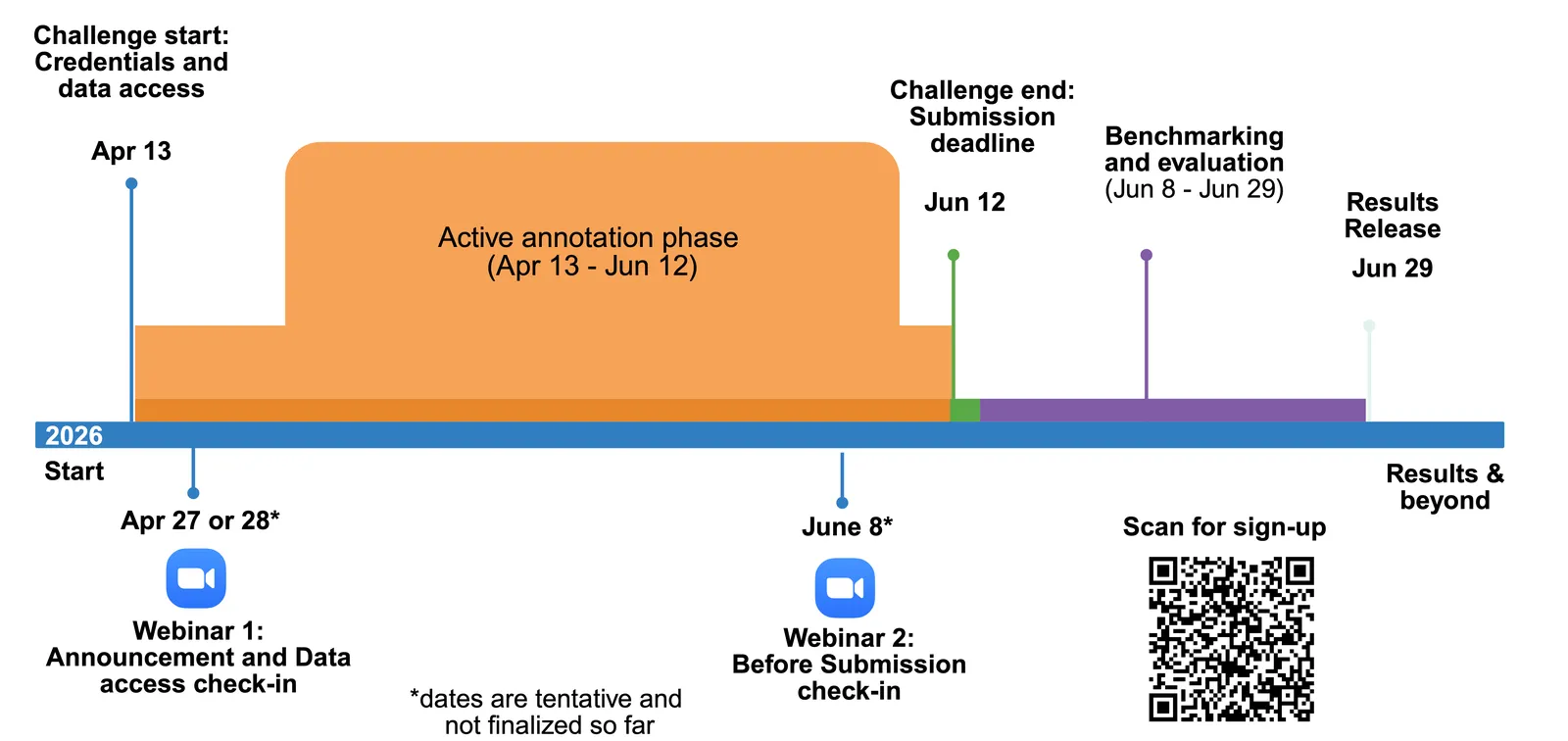
Figure 1: HIPC scRNA-seq Annotation Benchmark Challenge: Timeline, illustrating key milestones from data access distribution (April 13) through the active annotation phase, submission deadline (June 12), benchmarking and evaluation, and final results release (June 29).
1. Overview and Objectives
The primary goal of this benchmark is to evaluate the consistency, accuracy, and resolution of scRNA-seq cell-type annotations across independent teams and pipelines.
Instead of relying on a single absolute gold standard, we use a multi-criteria evaluation framework that reflects how annotations are generated and interpreted in real research settings. Specifically, we focus on three complementary dimensions:
Annotation Concordance: We assess how closely submitted labels align with the original study annotations, after mapping both to a shared Cell Ontology, and whether they are supported by established marker genes.
Inter-Team Consensus (reproducibility): We measure agreement across teams annotating the same datasets. Strong consensus suggests robustly identifiable populations, while disagreement highlights ambiguous or method-dependent cell states.
Granularity: We evaluate the level of resolution achieved by different pipelines, ranging from broad categories (e.g., T cells) to more specific functional or differentiation states (e.g., CD8⁺ effector memory T cells).
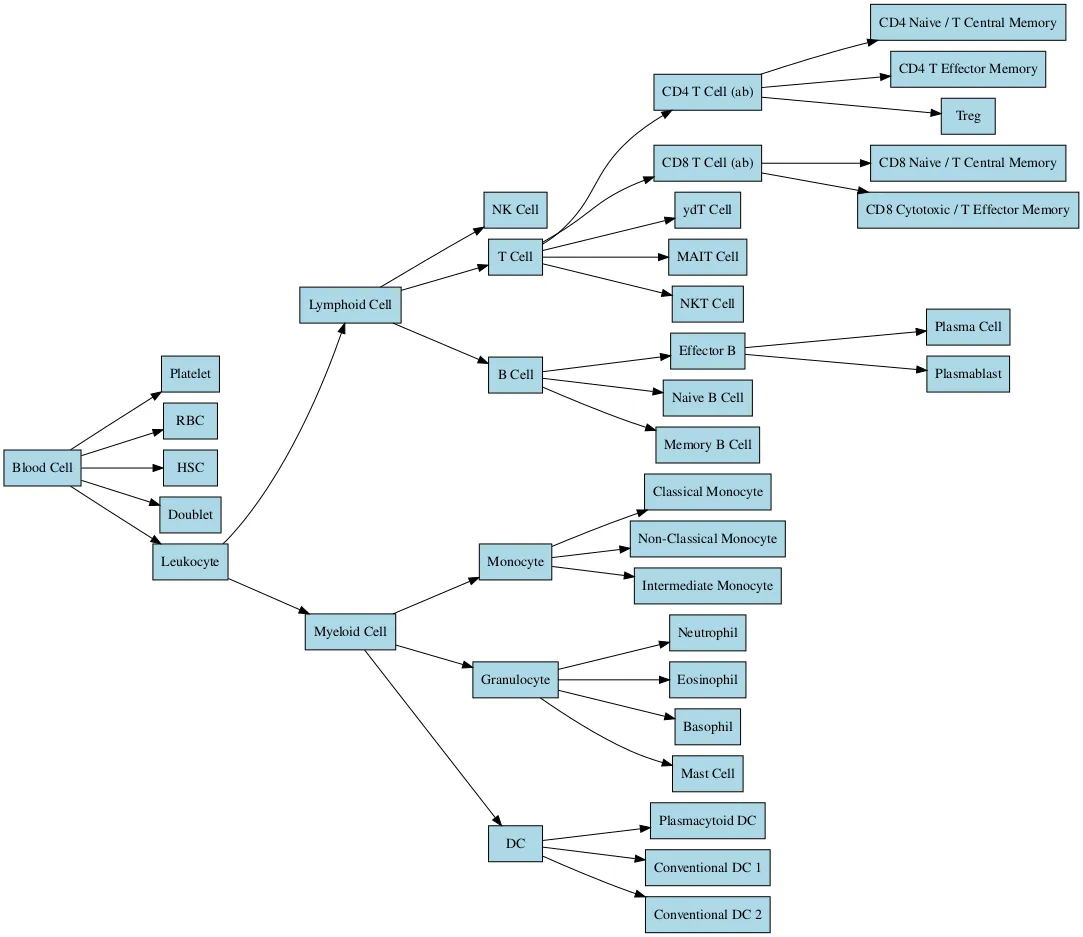
Figure 2: Single-cell transcriptomic cell type annotation hierarchy. Hierarchical classification of blood cell types identified by scRNA-seq. From the root (Blood Cell), cells are stratified into major lineages (Platelet, RBC, HSC, Doublet, and Leukocyte). Leukocytes are further resolved into Lymphoid (NK, T, and B cell subsets) and Myeloid (Monocytes, Granulocytes, and Dendritic Cell subsets) compartments.
2. Data Access
All datasets are distributed through a secure, private infrastructure to preserve the integrity of the benchmark.
- Each team is assigned a specific "recipe_data." After logging into the ImmuneSpace file server, you will only see the studies assigned to your team.
- Unique login credentials will be emailed to each team lead.
- Original study identifiers have been removed. Please refer only to the internal IDs provided (e.g., infection_study_01, vaccination_study_01).
Privacy Note: Dataset assignments may overlap across teams, but specific assignments are confidential. To ensure independent analyses, please do not discuss assigned studies with other participating groups.
3. Annotation Tasks
Participants are asked to annotate immune cell populations within their assigned scRNA-seq studies. To ensure fair comparison and reproducibility, please follow these guidelines:
- All labels must be mapped to the provided standardized Cell Ontology dictionary and match the nomenclature exactly (Refer to CT_Ontology_Spreadsheet_*.xlsx).
- Aim for the highest level of granularity you can support with confidence.
- Fully document your workflow on GitHub. Reproducible code (R Markdown or Jupyter Notebook) is required so that all submissions can be audited and compared transparently.
4. Submission Workflow
Participants will have ~ 2 months to complete the annotation process ().
Workflow overview:
- Download. Access your files at immunespace.org/annotation_benchmark/data_recipe_[number] using your Nginx credentials provided via email.
- Annotate: Perform manual or automated annotation. You must map all results to the provided standardized cell ontology dictionary (refer to celltype column in CT_Ontology_Spreadsheet_*.xlsx).
- Validate: Ensure your submission is in the required .tsv format and includes reproducible code.
- Submit: Upload your final bundle via the secure Google Form mentioned in Step 6.
5. Reference Materials
In your private data directory, you will find a Reference Folder containing:
- The Cell Ontology (.xls) file for label mapping. Use the celltype column as the basis for your predicted cell type labels.
- A Visual Guide provided by the HIPC processing team to assist in identifying expected populations.
- A "Fake Example" folder to demonstrate the correct file structure for your submission.
6. Final Step: Submit Results
Submission Requirements: Alongside the GitHub code repository link, each team must provide a single .zip folder containing the following:
| File Name | Content | Format/Requirements |
|---|---|---|
| annotation.tsv | Final cell labels | Must include cell_barcode, predicted_cell_type. Submit a separate file for each dataset annotation. |
| readme.docx | Methodology | Detailed methodology used software versions and any manual intervention steps. |
Ensure you have read the Submission Requirements and General Rules & Guidelines in full before submitting your entry.
For technical issues, including data access or cell ontology questions, please use following space in the ImmuneSpace Solution Center.